Digital Pathology integration
We can identify two basic phases in a Digital Pathology reference workflow:
Whole Slide Image (WSI) file and metadata ingestion into the archive
Open VIEW on a specific digital slide (or case) with an in-context call
Data ingestion
Actors:
Digital slide producer (e.g. the WSI scanner)
Metadata owner system (e.g. Laboratory Information System)
FEED
PACS
Note: we assume that there is an integration between the digital slide producer and the metadata owner system.
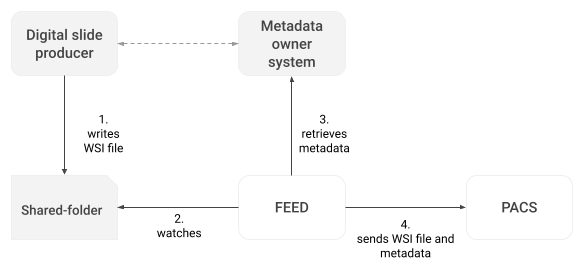
Data ingestion workflow
The digital slide producer writes the WSI file into a shared folder (e.g. NFS folder);
FEED sees that a new WSI file has been written in the folder;
FEED queries the metadata owner system for patient and case information;
FEED sends the WSI file and the metadata to the PACS component, where the digital slide and metadata are stored and indexed.
Metadata retrieval transaction
With this transaction, the FEED script will retrieve the metadata from the metadata owner system.
The request type will be an HTTP GET and will contain only one parameter, the slide barcode identifier, that can be derived from the name of the WSI file:
/service/path/metadata?slide=XXXXIn the GET request, it is possible to include a header with a pre-shared static API key as an authorization mechanism (e.g. Authorization: API-KEY).
The service should return the following data in JSON format:
Field | Type | Description |
|---|---|---|
| Required | Unique patient identifier (e.g. |
| Recommended | Patient’s name in the format |
| Recommended | Patient’s birth date in the format |
| Recommended |
|
| Required | LIS unique case identifier (max 16 characters) |
| Recommended | LIS unique identifier of the specimen |
| Recommended | Specimen display name (e.g. |
| Recommended | Specimen procedure or description (e.g. |
| Recommended | Topography (e.g. |
| Recommended | LIS unique identifier of the block |
| Recommended | Block display name (e.g. |
| Recommended | Block procedure or description (e.g. Displayed in the virtual tray |
| Required | LIS unique identifier of the slide |
| Recommended | Slide display name (e.g. |
| Recommended | Slide stain code (e.g. |
LIS response body example:
{
PatientID: "PID34125",
PatientName: "TURNER^KATIE",
PatientBirthDate: "19750902",
PatientSex: "F",
AccessionNumber: "24-H-00123",
SpecimenIdentifier: "S920939933800092655716259",
SpecimenAlias: "A",
SpecimenProcedure: "Breast Biopsy",
SpecimenBodySite: "BREAST",
BlockIdentifier: "S906723612258515086899",
BlockAlias: "A-1",
BlockProcedure: "Margin"
SlideIdentifier: "S899706197241433574521",
SlideAlias: "A-1-A",
SlideStainCode: "H&E"
}Virtual tray example and details:
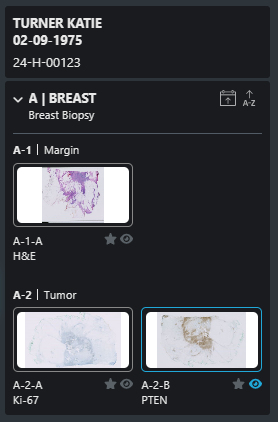
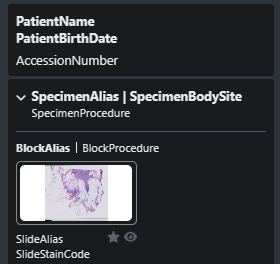
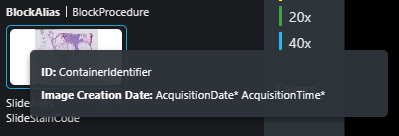
* provided by the FEED
WSI image availability callback
The PACS component can be configured to send an PACS webhooks | Image-Availability-Message to a third-party system to notify that the slides have been correctly imported into the system and are ready to be viewed.
For Digital Pathology, the image availability message will contain one series item for each digital slide, and the slide barcode identifier will be placed in the ContainerIdentifier.
The system will send a message after every new slide the archive receives, and the message will contain information about all the slides in the case.
VIEW integration
An external system like the LIS can open VIEW on a specific digital slide using the VIEW APIs | Access-Token-API-(GET).
In a typical use case, VIEW can be opened on a digital slide by specifying the accNum and containerIdentifier query parameters.
Additional integrations
VIEW allows additional integrations with third-party systems like the LIS.
Webhook events
VIEW notifies a third-party system every time a digital slide status changes. Details can be found in the VIEW webhooks section.
AI algorithms
It is possible to integrate third-party AI algorithms to receive and display findings on processed digital slides. This type of integration necessitates a project-specific analysis.
Snapshots
VIEW can send a snapshot of a digital slide taken by the user to a third-party system.
The viewer will forward a POST request to the service configured in the NgvConfiguration table under the name ApSnapshotUrl. If the field is empty, no message will be forwarded.
The request will contain the header Content-Type: application/json while its body will look like the following:
{
"username": "Administrator",
"timestamp": "2023-09-04T09:56:49.6213677",
"wholeslide":
{
"studyInstanceUID": "1.2.826.0.1.3680043.2.619.161.1027041362",
"seriesInstanceUID": "1.2.826.0.1.3680043.2.619.1612.1688546606484.1",
"accessionNumber": "I20-230705",
"containerIdentifier": "I20-230705-A-1-2"
},
"content":
{
"contentType": "image/jpeg",
"caption": "this is a caption",
"data": <base64 encoded image>,
"magnification": 5.434888974179961,
"region":
[
{
"x": 64332.641290268315,
"y": 6379.44867537442
},
{
"x": 71111.0686279783,
"y": 6379.44867537442
},
{
"x": 71111.0686279783,
"y": 8808.201141654868
},
{
"x": 64332.641290268315,
"y": 8808.201141654868
},
{
"x": 64332.641290268315,
"y": 6379.44867537442
}
]
}
}Focusing on the object content, it will contain:
the
contentTypeof the snapshot, always set toimage/jpeg;the
captioninserted by the user;the image exported in Base64 in the string
data;the zoom level (
magnification) at the time the image was exported;the coordinates (
region) relative to the exported image.
The remaining shared information is related to the session (username and timestamp) and case or slide identifiers.
